Phonemic SFE disconfirmed
« previous post | next post »
Last spring, I took a look ("Phonemic diversity decays 'out of Africa'?", 4/16/2011) at an interesting paper by Quentin Atkinson ("Phonemic Diversity Supports a Serial Founder Effect Model of Language Expansion from Africa", Science 4/15/2011). Atkinson argued that a survey of sound systems around the world supports the so-called serial founder effect (SFE) "in which successive population bottlenecks during range expansion progressively reduce diversity", just as a similar survey of human genetic and phenotypic diversity does. He also argued that the phonemic-diversity evidence points to an origin in Africa, again just like the genetic evidence.
I expressed some skepticism about this argument, mainly based on some of the choices that Atkinson made in quantifying "phonemic diversity". One choice that I considered in detail was the critical role played by a few features such as tone, which (on the time scale of human global migration) are at least as likely to result from innovation and areal spread as from survival.
Now Keith Hunley, Claire Bowern, and Meghan Healy ("Rejection of a serial founder effects model of genetic and linguistic coevolution", Proceedings of the Royal Society B, 2/1/2012) have taken another look at the genomic and phonemic predictions of the SFE. They chose a very different way of coding the distribution of phonemes — formally analogous to the way that they coded genetic variation — and this time, the phonemic data gave very different results.
Their abstract:
Recent genetic studies attribute the negative correlation between population genetic diversity and distance from Africa to a serial founder effects (SFE) evolutionary process. A recent linguistic study concluded that a similar decay in phoneme inventories in human languages was also the product of the SFE process. However, the SFE process makes additional predictions for patterns of neutral genetic diversity, both within and between groups, that have not yet been tested on phonemic data. In this study, we describe these predictions and test them on linguistic and genetic samples. The linguistic sample consists of 725 widespread languages, which together contain 908 distinct phonemes. The genetic sample consists of 614 autosomal microsatellite loci in 100 widespread populations. All aspects of the genetic pattern are consistent with the predictions of SFE. In contrast, most of the predictions of SFE are violated for the phonemic data. We show that phoneme inventories provide information about recent contacts between languages. However, because phonemes change rapidly, they cannot provide information about more ancient evolutionary processes.
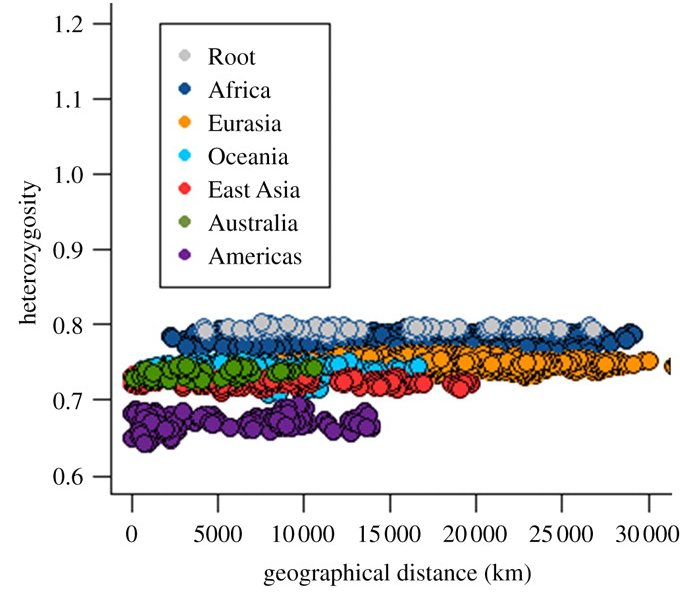
Here's their graph of between-population heretozygosity versus geographic distance, for their genetic data:
They note that
The top, grey-coloured points show the heterozygosity between the African San and the other 99 populations […]. The level of heterozygosity is roughly uniform whether populations are located nearby in Africa, or thousands of kilometres away. This uniformity reflects (i) a single African origin for all humans, (ii) a split between the population that would become the San and the founder of the remaining 99 populations, and (iii) relative subsequent isolation between these two groups.
The next tier of dark blue points shows the heterozygosity between the remaining 19 African populations and the 80 non-African populations. The level of heterozygosity is again uniform over thousands of kilometres. This uniformity reflects common ancestry for all non-African populations associated with an ancient out-of-Africa founder event. The remaining tiers are the product of subsequent splits and founder effects associated with the peopling of major geographical regions[…].
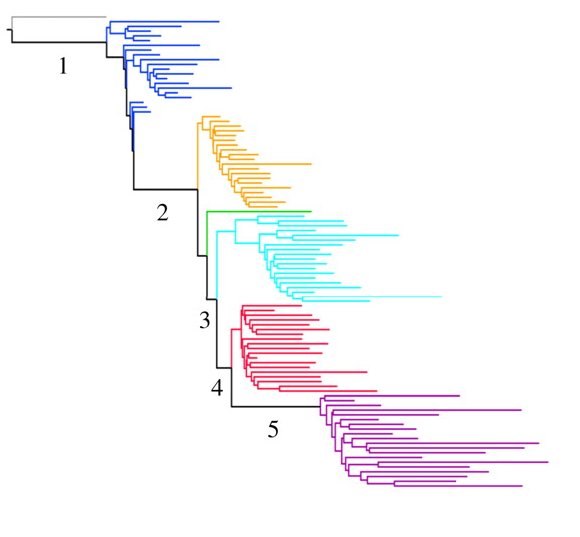
The corresponding rooted neighbor-joining tree:
They comment that
The test of treeness […] indicates that the tree is a good representation of the pattern of genetic variation, and that geographical distance explains little of the pattern of among-population genetic distance independent of the tree. All trees rooted on an African branch of the tree fit better than all trees rooted on a non-African branch of the tree. The best-fitting of all possible rooted trees separates the African San from the remaining 99 populations, and the level of variation within populations decreases steadily away from this node (signified by ever-increasing terminal branch lengths).
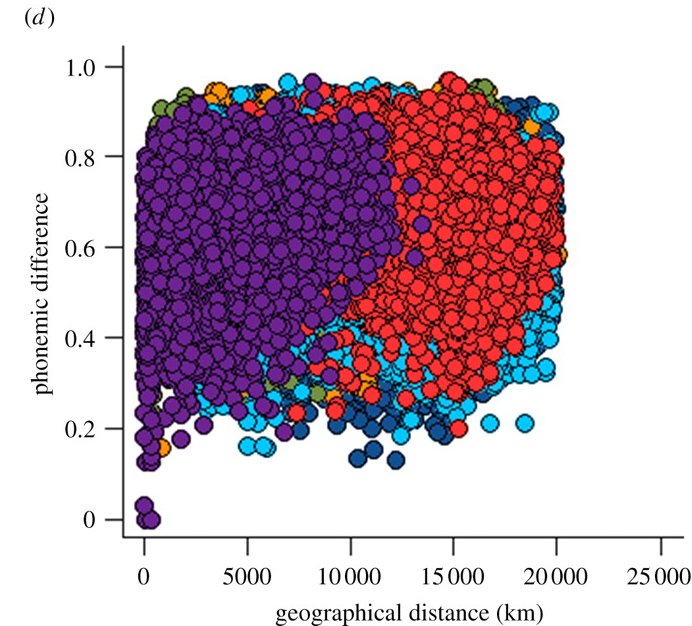
Here's their graph of between-language phonemic difference versus geographical distance:
They observe that
… the among-region pattern of phonemic variation is not tiered, but there is some evidence of a correlation between phonemic difference and the geographical distance in the plot. The correlation could be a by-product of SFE (i.e. it could reflect the tendency of phylogenetically related languages to be located near to one another). Alternatively, it could reflect sustained phonemic exchange (borrowing) between geographical neighbours, in which case it is inconsistent with SFE.
This pattern is not very consistent with any particular tree-structured hypothesis:
Several methods were used to construct a phoneme tree. Each produced a different topology; none were similar in topology to the microsatellite NJ tree and diagnostic output from each method strongly implies that phonemic variation is not tree-like. Figure 4c shows a midpoint-rooted phoneme tree produced using a Bayesian approach. Though there is some regional clustering, it contains considerably less geographical structure than the microsatellite NJ tree in figure 4a. Though the midpoint root does separate an African language from the remaining languages, African languages are dispersed throughout the tree.
In order to distinguish geographical patterns due to SFE from geographical patterns due to areal borrowing, they
… examined (i) the correlation between phonemic difference and geographical distance within the five best-sampled language families in our sample and within each geographical region, and (ii) partial correlations between phonemic difference and geographical distance within each region controlling language family membership.
Their overall conclusion is that "phonemic diversity has not been moulded at the global level by the same evolutionary processes that shaped neutral genetic diversity", and they note that these results are expected on common-sense grounds:
The genetic signature of founder effects persists in human populations in part because they accumulate variation slowly. In language, however, even if founder effects initially eliminated phonemes, rates of phonemic change are so high that the signal of loss would quickly disappear. While we reject the SFE process for phonemic data on both empirical and theoretical grounds, these data do provide information about recent contacts between languages.
And although their method of comparing phoneme inventories is arguably more appropriate than Atkinson's method of using a single numerical "diversity index", they argue that a comparison of allophones would be even better:
Phonemes are the sound categories that signal a difference in meaning between two words. For example, /d/ and /t/ are distinct phonemes in English because they contrast in the words <bad> and <bat>. But both /d/ and /t/ have a range of sub-phonemic allophones that are conditioned by both location within the word and non-linguistic demographic factors such as social class. For example, /d/ has voicing when it occurs between vowels, but it is partially or fully devoiced for most English speakers in word-final position. Variation in allophones is found in all languages and is a major driver of language change. In contrast, the level of phonemic variation within a language is small. Thus, if an SFE model does apply to language, it is more likely to affect allophonic variation. A daughter population would contain a subset of the allophonic diversity found in the parent, and the daughter would then be subject to processes of allophonic change, drift and selection that lead to sound change. Crucially, such changes are largely neutral with respect to phoneme inventory size. Unfortunately, there currently exist no databases of allophonic variation that would allow this hypothesis to be tested. In contrast, borrowing effects would be expected to be revealed in phonemic inventories as neighbouring languages converge on similar inventories due to contact.
I'm not entirely convinced by this argument. Genomic variation remains digital down to the level of single-nucleotide polymorphisms, but (as their phrase "partially or fully" hints) much allophonic variation doesn't naturally fall into qualitatively-distinct classes. Instead, we often need to describe such variation in terms of quantitative changes in complexly-conditioned distributions of continuously-valued measurements. As a result, the question of how to quantify comparisons becomes a complex one.
In the genomic case, we can count substitutions, deletions, translocations, etc. There may sometimes be questions about what "edit distance" to use, but I think (perhaps out of ignorance) that plausible, useful, and consensual answers are available. In the phonetic ("phonomic"?) case, we'd have to decide (for example) how to weigh X milliseconds of change in average aspiration duration in context A, against Y Hz of formant change in average vowel height in context B. (And after we settled that one, it would start to get really complicated…)
One plausible approach would be to define a metric in terms of the variances and covariances of the measurements involved, the relative frequency of the contexts, etc. But this would rely on a body of cross-language data that's REALLY different from anything that now exists.

dw said,
February 2, 2012 @ 9:58 am
Trying to measure the number of different allophones in a language strikes me as being about as difficult (and as useful) as trying to measure the number of different heights, or shades of skin color, in a population. The result will be almost entirely arbitrary, based on the granularity with which the measurements are distinguished.
[(myl) Their proposal, as I understand it, doesn't require "measuring the number of different allophones in a language", but rather depends on being able to quantify an "allophonic difference" between the speech patterns of two individuals or two speech communities. This would presumably have a qualitative part (e.g. does the language have final devoicing?) and a quantitative part (e.g. what is the distribution of voice onset time in initial unvoiced stops?).
In fact, there's an engineering problem (language identification, or "LID") for which one class of solutions involves placing the test sample in some sort of universal phonetic space, and then comparing its similarity to each of a set of languages for which training samples are available. The comparison metric can involve a "phonotactic model" which estimates the probability of different sequences of broad phonetic classes, or it can look at the distribution of quantitative measurements for certain kinds of sounds or sound sequences, or both.
The choices made are probably not the ones that you'd want to make to apply such ideas in the case under discussion, but the basic problems are similar.]
James Winters said,
February 2, 2012 @ 4:47 pm
"Crucially, such changes are largely neutral with respect to phoneme inventory size." — I'm not too convinced by this statement. If you lower the allophonic variation of a phoneme, then it lowers the functional load, meaning it's potentially more susceptible to merger (given a certain set of conditions: see Martinet, 1952). I would also add that if a phoneme frequency is only present in a certain number of constructions, especially if those constructions themselves are low in frequency, then the loss of speakers could, theoretically at least, see the loss of these phonemes without any merger taking place.
Joyce Melton said,
February 2, 2012 @ 7:07 pm
Phonemic choice in language may also be influenced by culture and environment. The whispers and clicks of the San people in their African deserts are admirably suited to the lifestyle of hunters in a quiet landscape where sound carries and is identifiable as to direction but hunters still need to communicate with their hunting partners. The reduced phoneme inventory and reduplicative languages of some Polynesian populations, on the other hand, are well suited to communicating in the middle of the swoosh and splash of surf, wind and tide. Does either situation have that much to do with the languages ancestors may have spoken?
Ran Ari-Gur said,
February 2, 2012 @ 7:10 pm
The initial proposal was so absurd and ill-considered that it's sad that such a formal debunking was necessary, especially one requiring so much skill and effort; but these debunkers have done a really impressive bit of work.
Bruce Lin said,
February 2, 2012 @ 10:31 pm
That Figure 4c spiral is beautiful. How come my field doesn't produce amazing graphs like that? I can imagine that it doesn't work well for certain sets of data though – the rays could start to collide with the circumferential arcs.
Rohan F said,
February 3, 2012 @ 6:22 am
@Joyce Melton, how would you go about testing that proposition? As a counterexample, compare the Bushman languages to those of the ancient hunter-gatherer populations of Australia. Plenty of Aboriginal languages were spoken in desert areas not dissimilar from the Kalahari, but most of them have pretty ordinary-sized phonemic inventories. Pitjantjatjara and Warlpiri, both still spoken in central Australia, have 17 and 18 consonants respectively and both have short and long versions of just 3 vowels. The largest Australian inventory I know of is Aranda, which has 52 consonants and two phonemic vowels, but its inventory is large only because all of its 26 primarily articulated consonants have a corresponding phonemically labialised form.
Bob Ladd said,
February 3, 2012 @ 6:56 am
@Rohan F: Thanks for the Australian counterexample. I've always been skeptical of the claim that clicks are somehow adaptive in the Southern African hunter-gatherer context, and the Australian case at least shows that clicks don't necessarily arise in a similar culture in a similar environment.
As for Joyce's doubts about the relevance of what the ancestors spoke, consider another evolutionary speculation according to which it's not a coincidence that you find retroflex stops – which are unusual in the world's languages – in India and Australia. The idea is that the peopling of Australia happened via a S/SE Asian route so that Dravidian and Australian retroflex sounds go back to the speech of some common ancestor. If I have to choose between Just So Stories, I find that one more compelling (and potentially more testable on the basis of genetic and archaeological data) than the suggestion that clicks arose to facilitate cooperative hunting in Southern Africa.
Richard Sproat said,
February 3, 2012 @ 7:57 am
Worth pointing out for those who haven't seen it that volume 15 of the journal "Linguistic Typology" has a large number of papers on Atkinson's article in "Science", along with a reply from Atkinson.
Those, combined with this new article, make for a rather large collection of papers devoted to this rather curious idea, whose main attraction seemed to be its popular-science-pressworthiness.
It will be interesting to see if the popular science press that promoted Atkinson's original paper will be as ready to promote this new paper.
Claire Bowern said,
February 3, 2012 @ 9:45 am
Thanks for the feedback everyone.
First, regarding allophone measurement: we agree that it would be very difficult, for the reasons that Mark mentions (and some of which I think we mentioned in the paper). Given that allophonic variation is contextual (depends both on the phonemic environment in the word and on the social context of the utterance), that makes it very difficult to equate to alleles — after all, alleles don't vary depending on who the individual is talking to! Our point here though was more that phoneme are not a good equivalent to alleles.
It may be true that lowering the allophonic variation in a phoneme lowers its functional load (though I doubt that in the general case). And while lowering functional load may well lead increased likelihood of merger, that effect probably gets washed out once you consider that greater positional allophonic variation also leads to increases in likelihood of partial mergers. But these are empirical questions that could be tested with a decent sound change database. Unfortunately constructing such a database is only marginally less difficult than constructing an allophone database.
Regarding Figure 4c – it was made with FigTree, though there are other programs which can do that.
Regarding Australia: Most Pama-Nyungan have retroflex consonants because they were preserved from Proto-Pama-Nyungan. We should note, however, that the primary articulation for these consonants in Australia is with tongue retraction rather than with "proper" retroflection – that is, the contact tends to be apical, not subapical. However, one of my beefs with the literature on Australian phonetics is an unwarranted assumption of uniformity – just because the phoneme inventories are similar, it doesn't follow that all the phonetic detail is uniform too; but that's another story.
Finally, regarding press; a few news organisations were interested in the initial pitch, but lost interest when they realised that we didn't have a good story here about human origins.
Joyce Melton said,
February 4, 2012 @ 3:07 am
@Rohan F, I'm not sure it's testable in that way. Are there any other language groups who have lived as long as the San have in the same desert area? Too many variables, I guess. But I have been struck by the observation that the San language is almost indistinguishable at first hearing from natural sounds of a desert like the one I grew up in.
Perhaps the Australians moved too often? Perhaps some other adaptation served them as well? Like I said, too many variables.
And too, the only two language groups I've been able to come up with that fit this hypothesis are the San and the Polynesians.
Chris Davis said,
February 5, 2012 @ 4:02 am
@Joyce Melton: I should think that would be a good reason to abandon the hypothesis.
Jess Tauber said,
February 6, 2012 @ 1:51 am
Is anything in this hypothesis linked to recent findings of genetic hybridization between Homo sapiens and Neanderthals, for ex-Africa populations to the west, and Denisovans to the east? In gibbons or siamangs (can't remember the specifics) such hybridization between subspecies results in altered call structures. Certainly if there was the linguistic analogue of such an effect in the hybrid human populations, that would throw a monkey wrench (pardon the pun) into the model.
Rory Van Tuyl said,
February 9, 2012 @ 4:02 pm
Science Magazine has, at long last, published three "Technical Comments" in response to Atkinson's original article. All are unstinting in their criticism of his paper "“Phonemic Diversity Supports a Serial Founder Effect Model of Language Expansion from Africa” Science 15 April 2011, p. 346.
To view these detailed critiques, go to:
http://www.sciencemag.org/content/current#TechnicalComments
Joyce Melton said,
February 12, 2012 @ 1:20 am
@Bob Ladd. I don't think I suggested "that clicks arose to facilitate cooperative hunting in Southern Africa". I did suggest that a click language was well-suited to that occupation in that place and the implication could be made. Personally, I think clicks arose by random fluctuation in language and that they may have survived because of their suitability in that situation.
That would deal with their absence in other similar areas, clicks do seem to be at an extreme of language variation and they may just have not happened elsewhere. Nor did they travel much.
Reduplicative speech patterns, on the other hand, seem much more likely to have arisen by chance. But I'm not a language researcher, just an interested amateur.
Jess Tauber said,
February 17, 2012 @ 6:36 pm
Some years ago I did a phonosemantic study of one click language, based on a newly published dictionary that captured more phonemic contrasts. I found that there was a clear internal structure to roots, but not in general the kind of things one sees, say, in ideophones. Instead, what I saw was more in line with compacted morphological structure. The click-blocks had a strong link to the notion of hidden, difficult, or restricted moisture resources- which in the context of the desert environment in which the speakers make a living, makes some sense. So is this a holdover from some ancestral population predating the split, genetically, between themselves and the other linguistic lineages (implying we all hail from the hot place), or has it developed more recently as the environment they lived in changed, or was migrated into?
Phonemic Diversity and Vanishing Phonemes: Looking for Alternative Hypotheses | Replicated Typo said,
February 20, 2012 @ 7:38 am
[…] of a serial founder effect model of genetic and linguistic coevolution (Language Log provided some good coverage on this). To continue with the pile-on Science published three technical comments (see here, here […]
Florian Jaeger said,
March 3, 2012 @ 7:00 pm
And here is one more technical comment on Atkinson that just came out in Science. We conduct several large scale statistical simulations to assess the Type I error rate of Atkinson's analysis: http://www.sciencemag.org/content/335/6072/1042.1.full.pdf
The meat is, of course, in the supplementary materials (http://www.sciencemag.org/content/335/6072/1042.1/suppl/DC1). We hope that this approach to assess the support for hypotheses about typological distributions (incl. hypothesis about the origin of language based on typological distributions) will be of use to other researchers.
Using tools from evolutionary biology in cultural evolution | Replicated Typo said,
March 6, 2012 @ 1:38 pm
[…] (2011) paper on a serial founder effect is criticised (although not as tough as some other responses), but the authors suggest that "Regardless of the outcome of this debate, Atkinson’s […]