'Cause after all, he's just a vasopressin receptor
« previous post | next post »
To a recent article about metamaterials ("Chinese boffins crack invisible-shed problem", 9/3/2008), Lewis Page at The Register added this disclaimer:
WARNING: Habitual use of our science coverage may harm your baby and others around you. Such use has been linked to incidence of the King's Evil and is thought to contribute to global warming, entropy, substance abuse, terrorism, hair loss, volcanoes, orbital decay, intellectual languor and decline of moral standards in team sport.
Unfortunately, this warning is not yet required by the FCC. But if you're in the market for some entropy, intellectual languor and decline of moral standards in team sport, it'll be hard to top the past week's reports about the "monogamy gene" (AKA the "infidelity gene", the "divorce gene", etc.). A small sample of the headlines (with linked stories):
"Is monogamy genetic?"; "Baby, my genes made me do it"; "Some men carry 'commitment-phobia' gene"; "Is Lover Boy a Louse? It May Be Genetic"; "Infidelity: It's All In the The Genes"; "Would you abort George Clooney?"; "Study: For men, genetics might untie marital bonds"; "Marriage Woes? Husband's Genes May Be At Fault"; "Marriage problems? Husband's genes may be to blame"; "Study Finds Fear of Commitment May Be in a Man's Genes"; "Commitment phobes can blame genes: A man's reluctance to marry may be down to a genetic 'flaw', say researchers"; "Marital crisis? Blame it on male genes"; "'Bonding Gene' Could Help Men Stay Married"; "Divorce gene linked to relationship troubles"; "Scientists Discover the Monogamy Gene"; "Gene Variant Holds The Key To A Long And Happy Marriage"; etc., etc.
You can pretty much guess how this is going to turn out. The researchers found a small effect — the biggest result was a 5% difference in group means on a "Partner Bonding Scale", for 41 (out of 1,104) guys who were homozygotic for one of the 11 alleles examined at one of three loci. There was a lot of overlap in the distributions — the biggest effect size was about 0.38 — and most of the differences and effect sizes were much smaller. And because it was a twin study, the small group of subjects with the genetic pattern associated with the biggest difference was effectively even smaller, and may well have shared a large number of other cultural and genetic traits.
But most of the media coverage presented this as another triumph of contemporary genetic Calvinism: the fate of our relationships is written at birth in the book of our genes.
The stories were not nearly as bad as the headlines, for the most part, but I'm sure that most readers not innoculated against MSM science reporting will have come away from the experience with a stiff dose of misinformation. Thus the Washington Post's lede was
Whether a man has one type of gene versus another could help decide whether he's good "husband material," a new study suggests.
A study of Swedish twin brothers found that differences in a gene modulating the hormone vasopressin were strongly tied to how well each man fared in marriage.
And the LA Times went that extra distance:
Forget the prenup. Couples pondering whether to tie the knot should proceed straight to gene testing.
I'm happy to say that the background of this work is some first-rate science — a fascinating body of research about the neurochemistry of pair-bonding in monogamous and non-monogamous species of vole. At least, *I* think it's intriguing stuff, and so I'm motivated to explain it; and the recent (much weaker) human study doesn't make any sense at all without this background; so if you care about the neurochemistry of love, or about the rhetoric of science journalism, or both, just pour yourself a cup of coffee, find a comfortable seat, and listen up.
Among the many scientific papers on this subject, an especially good review can be found in Larry Young and Zuoxin Wang, "The neurobiology of pair bonding", Nature Neuroscience 7(10): 1048-1054, 2004.
The basic idea is that pair-bonding, at least in these rodents, is a form of conditioned-response learning, mediated by "the interaction of the ocytocin, AVP [arginine vasopressin] and dopamine systems within the reward circuitry". The part of the story that we care about today focuses on a brain receptor for AVP, which plays a critical role in the ability of rodents to recognize one another as individuals:
Selective V1aR [vasopressin 1a receptor] antagonist … administered into the lateral septum of rats … inhibits social recognition, whereas infusion of AVP or overexpression of the V1aR in this region enhances social recognition abilities. V1aR knockout mice also exhibit a complete loss of social recognition. However, … V1aR knockout mice perform normally on other olfactory and cognitive tasks, suggesting that this deficit is specific for social discrimination.
Thus "[g]iven this role of oxytocin and AVP in social recognition and the interaction of these peptides with mesolimbic dopamine, a reasonable hypothesis is that pair bonding results from the convergence of social discrimination circuits and the reinforcing properties of the mesolimbic dopamine reward circuit."
To become monogamous, in other words, a male rodent needs to be able to recognize his mate, and to allow mating with her to drive the reward and motivation system of reinforcement learning. If this works, as the song says, he becomes "accustomed to her face" — or maybe her smell, but whatever — and he starts to finds her company intrinsically rewarding. He's hooked; in effect, he's addicted to her. (There's a parallel process on the female side, but today we're talking about males.)
Voles have played a special role in this research, because prairie voles are monogamous and biparental, but the (closely related) montane and meadow voles are not. One of the genetic bases for this difference in behavior seems to be located in the AVPR1a gene, where the species differ in the length of a "microsatellite" sequence about 660 base pairs upstream of the transcription start site. This difference seems to result in brain-region-specific differences in the distribution of vasopressin receptor 1a — prairie voles have more of these receptors in the ventral pallidum.
And just by manipulating this genetic sequence, you can make those casual-sex meadow voles more like their pair-bonding prairie vole cousins — or at least you can make them spend more time hanging around with their partners after sex:
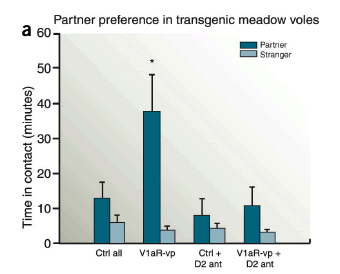
Male meadow voles overexpressing the V1aR receptor in the ventral pallidum (V1aR-vp) showed enhanced mating-induced partner preferences compared to control animals (Ctrl all). Infusion of a D2 receptor Antagoist (D2-ant) before mating abolished the partner preference in these males (V1aR-vp + D2-ant).
"Partner preference" is measured in an apparatus that
… consists of three chambers connected by tubes. The 'partner' and a novel 'stranger' are tethered in their own chambers, whereas the subject is free to move throughout the apparatus during a 3-h test. Pair bonding is inferred when subjects spend significantly more time in close proximity to the partner compared to the stranger (partner preference). In prairie voles, mating facilitates formation of partner preference…
In other words, after two prairie voles hook up, the male would rather hang out with the female he's just had sex with, compared to a female he's never met. Normal meadow vole males prefer the stranger, or would rather just be alone; but "transgenic" meadow voles, with more V1aR receptors in their ventral pallidum, act more like prairie voles in this test — unless they've been treated with a dopamine antagonist.
Against this background, it was quite reasonable for a group of researchers to look at the analogous genetic effects in humans.
Now, let's note in passing that we shouldn't really hope to find a "monogamy gene" or an "infidelity gene" or whatever on the basis of the vole research, since the species difference (at least the one that's been studied) involves a genomic variant whose effect on pair-bonding is apparently to help an animal tell other animals apart. (Like, how can you bond with a female if you can't recognize her when you see her again?)
And it's not clear that human males with commitment issues have any particular problems telling women apart — or that AVPR1a variation in humans is associated with other similar social-interaction problems. For example, Wassink et al., "Examination of AVPR1a as an autism susceptibility gene", Molecular Psychiatry 9:968-972, 2004, "did not demonstrate a disease-causing variant in the coding sequence", and concluded that "differences in AVPR1a at the amino-acid level are unlikely to confer genetic vulnerability to autism". (Though there *are* possible associations between AVPR1a and other personality traits that might have an impact on relationships: creative dance performance, age at first sexual intercourse, musical memory, balance between short- and long-term thinking, etc.)
Anyhow, genotyping has gotten to be cheap, and relationship problems are important, so it's well worth a shot. And the study responsible for all those headlines was published as Hasse Walum, Lars Westberg, Susanne Henningsson, Jenae M. Neiderhiser, David Reiss, Wilmar Igl, Jody M. Ganiban, Erica L. Spotts, Nancy L. Pedersen, Elias Eriksson, and Paul Lichtenstein, "Genetic variation in the vasopressin receptor 1a gene (AVPR1A) associates with pair-bonding behavior in humans", PNAS published online 9/2/2008. The abstract begins:
Pair-bonding has been suggested to be a critical factor in the evolutionary development of the social brain. The brain neuropeptide arginine vasopressin (AVP) exerts an important influence on pair-bonding behavior in voles. There is a strong association between a polymorphic repeat sequence in the 5' flanking region of the gene (avpr1a) encoding one of the AVP receptor subtypes (V1aR), and proneness for monogamous behavior in males of this species. It is not yet known whether similar mechanisms are important also for human pair-bonding.
However, there's no homologous genomic variation in humans:
Human AVPR1A is situated on chromosome 12q14–15.. Whereas there is no sequence in the human AVPR1A 5' flanking region homologous to the one found in prairie voles, humans do have three repetitive sequences in this region that are polymorphic: A (GT)25 dinucleotide repeat, a complex (CT)4-TT-(CT)8- (GT)24 repeat (RS3), and a (GATA)14 tetranucleotide repeat (RS1). […]
The aim of this study was to investigate whether variability in the 5' flanking region of AVPR1A affects pair-bonding behavior in humans as it does in prairie voles. To this end, the three repeat polymorphisms of the AVPR1A were genotyped in adult men and women from the Twin and Offspring Study in Sweden (TOSS), comprised of 552 same-sex twin-pairs and their spouses/partners. All subjects were assessed with respect to various indices of the quality of the marital relationship, including a new scale–the Partner Bonding Scale (PBS)–which is comprised of items that correspond to the behavioral patterns observed when measuring features of pair-bonds among nonhuman primates.
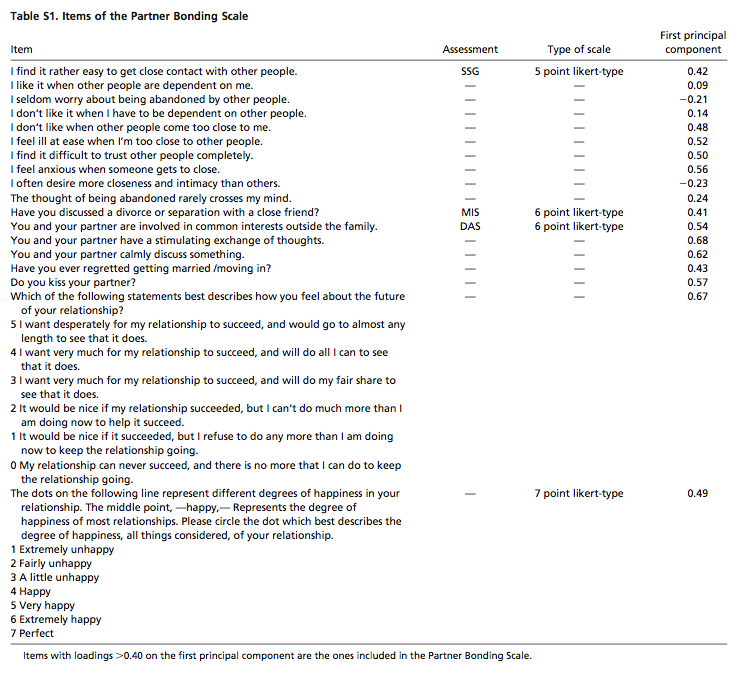
Rather than measuring whether subjects prefer to hang with their partners after sex or with strangers, as in the animal studies, Walum et al. devised a questionnaire whose responses they could turn into a "Partner Bonding Scale". The questions that they tried are below 13 (?) of them were used to calculate a Partner Bonding Score (PBS), which among their subjects ranged from 5 to 66:
(Click on the table for a larger version.)
However, it turned out that this "Partner Bonding Score" measure did not correlate especially strongly with the genomic variations in their sample. Only one of the alleles (RS3 allele 334) really stood out as especially different from the others — and the mean PBS score for this group was 46.2, compared to mean PBS scores from 45.6 to 48.8 for other alleles:
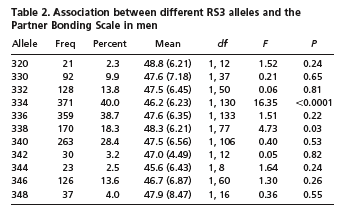
Why is allele 334 different from the other alleles P < 0.0001? Mostly because it was common enough for a small difference in mean values to be very unlikely to have arisen by chance. This is yet another example of the distinction between statistical significance and practical significance.
However, the effect of allele 334 on PBS was increased by looking at those men who were homozygotic for this allele (i.e. had two copies), rather than those who had one copy combined with one copy of another allele (i.e. were heterozygotic), compared to the men with no 334 allele. The guys with two copies of 334 had PBS of 45.5 (sd=6.71), compared to PBS of 48 (sd=6.5) for guys with no 334 at all. That's a mean difference of about five percent, and an effect size of 0.378, which is a moderate-sized effect, though not a spectacular one.
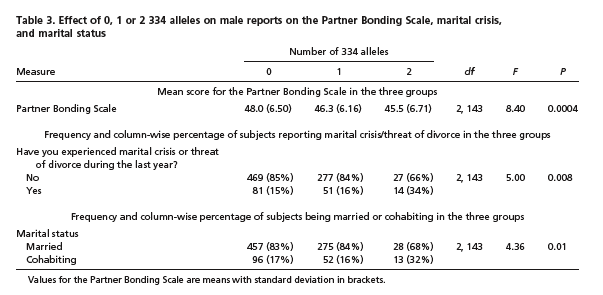
And the allele-334 homozygotes were quite a bit more likely (in percentage terms) to report a "marital crisis" within the past year (34% compared to 15%), and quite a bit less likely to be married rather than cohabiting (68% vs. 83%):
However, we ought to be worried that there were only 41 allele-334 homozygotes in this study, so that the confidence intervals for these percentages are going to be fairly broad. And more seriously, there might well have been some non-sampling error, say because (some of) those 41 guys might share some significant non-genetic characteristics — maybe they were mostly immigrants from a particular country, or fisherman from a particular region, or something like that. In any case, this was a twin study (only, I think, by the accidental fact that a twin study was available for the researchers to piggy-back on), so that many of the 41 334-allele homozygotes were probably pairs of twins, who presumably shared many familial and cultural features as well as other genetic features.
Note that the 41 guys with two copies of 334 only had about 7 more instances of marital crisis, and about 6 fewer marriages, than would be expected given the proportions in the other groups — so it wouldn't take many participants with shared socio-cultural (or other correlated genetic) characteristics to create this effect.
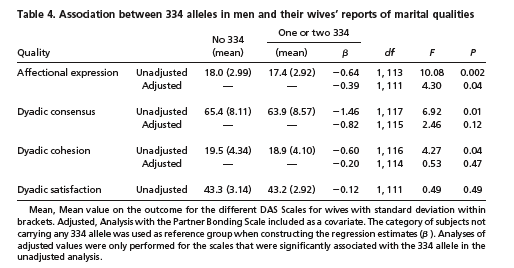
The last issue has to do with how the men's partners perceived the relationship:
… we investigated whether the genotype of the men influenced marital quality as perceived by their spouses. For this purpose, we used the Dyadic Consensus, Dyadic Satisfaction, Dyadic Cohesion, and Affectional Expression subscales from the Dyadic Adjustment Scale (DAS) (19), measures frequently used to evaluate the quality of marital relationships.
Thus there was no significant effect on "Dyadic satisfaction" — the partners of the allele-322 guys were just as happy with the relationship, overall, as partners of other men were. There were small but statistically significant differences in "Affectional expression" (3% difference in means, effect size d=0.20), "Dyadic consensus" (2.3% difference in means, effect size d=0.18), and "Dyadic cohesion" (3% difference in means, effect size d=0.14):
This last difference (3% difference in group means, effect size of 0.14) is what the BBC's story describes as
"Men with the 334 version of the AVPR1A gene earned lower scores from their partner/wife for strength of relationship bond."
If you know what "effect size" means, you'll recognize that this can be translated as
"If you pick a random man with the 334 allele of the AVPR1a gene and a random man with a different allele, the 334 guy will get a higher score from his partner for strength of relationship bond about 47% of the time, and a lower score about 53% of the time."
My conclusions:
The vole pair-bonding research is terrific stuff, which leaves a lot of fascinating open questions. The human study probably means something, though the effect sizes are small enough that the idea of pre-nup genotyping is preposterous. More seriously, it seems far from clear to me that the results have anything essential to do with the vole pair-bonding research — at least, if the Young/Wang model for the role of AVPR1a in vole pair-bonding is correct, then the basic mechanisms must be entirely different.
(See here and here for some other relevant things that AVPR1a might be doing in humans…)
And it wouldn't surprise me to learn that you could get similar effects on the authors' "Pair Bonding Score" from similar studies of alleles of a wide variety of other brain hormone receptors, which presumably correlate in a similarly weak way with various personality traits that influence relationship success.
TW said,
September 5, 2008 @ 9:02 am
Excellent take on the misinformation and bad reasoning in a lot of science reporting today. What's even more interesting is how these memes take on a life of their own and come to dominate the way that reporters-and ultimately the public at large-interpret events. Some anthropologist needs to look at western society's cargo-cult relationship with science and scientists using this MSM reporting as primary data.
Ryan Denzer-King said,
September 5, 2008 @ 12:04 pm
The focus of science reporting on exaggerating claims of genetic predisposition raises an interesting point: why do people like to hear this? Is it somehow comforting to be told that we are not responsible for our actions, that we are merely biological machines plodding through a predetermined courses?
I find it neither comforting nor likely.
Tom Duff said,
September 5, 2008 @ 3:21 pm
I think what's happening here is that people today look to science for moral guidance where they once looked to fables. The researcher replaces the wise grandfather telling tales by the fire, and three voles in a maze replace Br'er Rabbit in his briar patch.
A striking thing here is that the moral that the human researchers take from the vole study is nearly as abstract as what the science reporters draw from the human research, so by the time it gets to the general public the excellent results of the vole research are completely denatured.
BTW, I nominate "You can pretty much guess how this is going to turn out" for the official motto of the Language Log science journalism watchdog desk.
Mark Liberman said,
September 5, 2008 @ 5:27 pm
Tom Duff: I think what's happening here is that people today look to science for moral guidance where they once looked to fables. The researcher replaces the wise grandfather telling tales by the fire, and three voles in a maze replace Br'er Rabbit in his briar patch.
I think this is exactly right. In fact, I once suggested a somewhat different version of the same idea — "
Biblescience stories", 12/2/2006:Tom's analogy between Br'er Rabbit in his briar and the voles in their three chambers is a beautiful one.
john riemann soong said,
September 7, 2008 @ 1:19 am
I find it disturbing whenever subject-answered surveys are used in cognitive science to try to turn things into a "scale". It reminds me of the Spanish/English masculine-feminine linguistic-cultural study, or whatever that was.
"But most of the media coverage presented this as another triumph of contemporary genetic Calvinism: the fate of our relationships is written at birth in the book of our genes."
The problem I think is the public not even realising what role genes really play. It's the public perception caused by knowing that there are genes that make fruit last longer, eels glow in the dark or whatnot, not realising that the mechanisms are much more subtle. The whole analogy as "genes as blueprints" has been quite harmful I think.
I am struck by the fact that genes are often surprisingly deterministic of personality, as identical twin studies sometime show. There are no genes that code for having the tendency to sneeze in elevators on purpose, but gene *complexes* can do that. The public has been presented with the butterfly effect, but can someone SOON give the impression to the public that that applies for genes?
Mark Liberman said,
September 7, 2008 @ 7:12 am
john riemann soong: I find it disturbing whenever subject-answered surveys are used in cognitive science to try to turn things into a "scale".
This practice started with IQ tests and other factor-analysis-based test instruments. These days, it's used by the psychometricians who design all sorts of aptitude and achievement tests; and also by social psychologists and clinicians intereted in measures of personality characteristics, emotional states and so on.
It's usually a serious error to take the reification of these scales seriously, as Cosma Shalizi explains here.
But when you need a metric (for a fair ranking of applicants to some educational program, or for a study of the efficacy of treatments for depression, or for this kind of study of the genetics of human pair bonding), what else are you going to do?
Promoting Good Coverage of Science in the Media « everyONE - the PLoS ONE community blog said,
March 26, 2009 @ 2:48 pm
[…] Log (a linguistics/language-related blog, which often covers topics like the semantics of science news headlines and the use of “mendacity […]